MCG researchers leverage the power of IBM’s Watson to improve targeted therapies for cancer
IBM’s Watson has beat human champions on Jeopardy.
Now researchers are hoping this icon of artificial intelligence will help people with cancer win as well by providing a rapid, comprehensive report of the genetic mutations at the root of their specific disease and the therapies that target them.
“We have to change our whole behavior in looking at tumors. We are missing too much, and too often treatment does not work for patients,” says Dr. Ravindra Kolhe, breast and molecular pathologist and director of the Georgia Esoteric & Molecular Labs LLC in the Department of Pathology at the Medical College of Georgia.
“With Watson, all oncogenes become targets.”
Today, for example, a pathologist would use polymerase chain reaction, or PCR, to analyze a melanoma biopsy for a single variation of the BRAF gene, which is present in about 50 percent of melanomas and results in production of an altered protein that helps the cancer grow.
There are drugs that directly target that mutated gene, but if the pathologist doesn’t find that variation, which happens in about 50 percent of patients, the patient will receive a more broad-spectrum chemotherapy regimen. “We know that there is often more than one gene involved, even when we find BRAF,” he says.
Watson has the ability to not only rapidly identify multiple variations in BRAF, but variations in hundreds of other genes known to contribute to the most deadly skin cancer, as well as the therapies to target them, Kolhe says.
“The majority of the time, we just tell patients they have a cancer,” Kolhe says. “Watson can help us provide more comprehensive, personalized care to patients.”
The young pathologist first worked with Illumina, a California-based biotech company that makes and markets systems to analyze genetic variations and biological functions, to validate the TruSight Tumor 170 panel, which helps give Watson a much broader cancer genomics perspective.
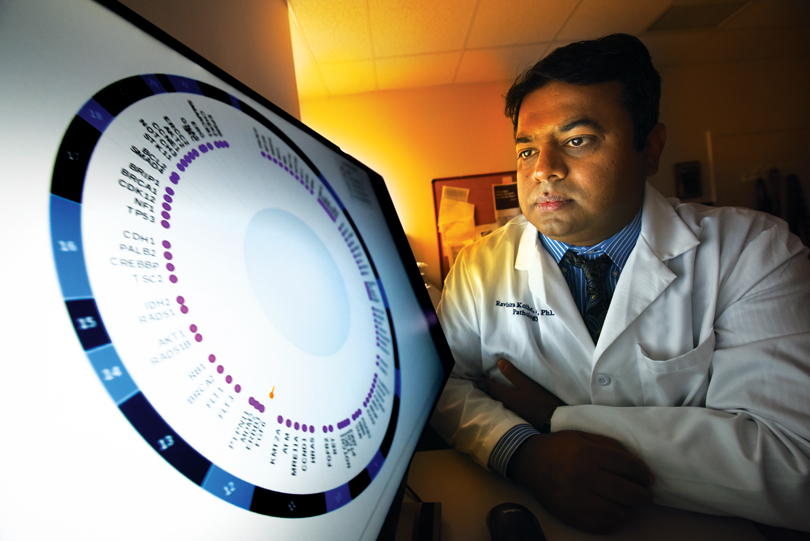

The panel contains known mutations on 170 genes commonly altered in cancers – mutations that, like those with BRAF, can transform previously inactive genes into oncogenes in eight common cancers, including lung, colon, breast, prostate as well as melanoma. Many of these genes are a player in multiple cancer types, including those not currently on the list. “In probably 90 percent of tumors you will find one or a combination of these 170 genes involved,” Kolhe says.
It also enables Watson to read all amino acids in the genes since it’s the amino acid changes that mutate the genes. It’s looking for dramatic change that might help make cells “immortal” and cancerous, Kolhe says. “One gene has virtually hundreds and hundreds of different combinations of amino acids and each amino acid in each combination can have mutations that are cancer causing.”
The panel also enables identification of gene fusions. Genes don’t normally fuse, and this unhealthy merger is another way cancer happens. Basically the promoter from one gene that is normally always turned “on” decides to also turn on another gene that is normally silent, as most of our genes are, Kolhe says.
Today’s standard for detecting fusions alone is much more limiting, he says. Fluorescence in situ hybridization can only examine one gene at a time and may not be able to determine which specific genes fused and so how best to target them.
“It’s a good but extremely limited tool,” Kolhe says, that sometimes provides no insight on the cancer. “That is one of the frustrating parts about pathology. You can tell by looking that you have a cancer, but we don’t know the oncogenesis or the way the malignancy happened,” he adds, particularly when you are looking one gene at a time.
“This is a leap in technology. Here we are looking at 170 genes and all the amino acids and all the fusions at one time.”
The exploration for a better way began about five years ago.
“We also started with a single gene in a single tumor but went one step further and decided to look at a group of genes in a single tumor,” Kolhe says. “Then we thought there might be some melanomas that are going to have BRAC1, a common breast cancer gene. There might be some melanomas that also have PTEN, a gene that is normally a tumor suppressor but when altered can become a player in many cancer types. Why not collect all the genes in all the tumors and design one panel and see if we can find something that is more comprehensive?”
Now Kolhe sequences DNA and RNA from a patient’s tumor and feeds the information to Watson, which, unlike traditional search engines, can gather evidence and analyze data but also generate knowledge.
“In about 20 minutes, Watson looks at what is abnormal in the sample, then takes the abnormalities and looks all around the world to see what are the drugs already used against them, what studies are underway against them, even other drugs out there that might be useful,” Kolhe says.
He likens the significantly broadened perspective to the difference between looking down through a traditional microscope and up through a Hubble space telescope. And it’s fast.
This type of comprehensive report might take 10 people 10 days to generate, if ever, Kolhe says.
Decisions about treatment clearly are still in the hands of physicians and patients, where they belong, Kolhe says, but Watson can make unprecedented information immediately available to make those decisions.
It also is helping identify potential new cancer-causing genes. As an example, during one of many test runs, Watson provided Kolhe a 30-page assessment of a patient with a rare soft tissue sarcoma, which included mutations on seven genes, where the mutations are, what they are known to do and options for targeting them.
There also was a list of gene alterations that have no known therapies, as well as gene mutations that were present but their contribution to disease isn’t currently known.
It is those unknowns Watson finds that might help researchers like Kolhe also identify new genetic contributors – and treatment targets – in a variety of cancers and make Watson a valuable scientific tool as well.
In the earlier stages, Kolhe’s lab team and the IBM team were actually doing the labor-intensive work of manually combing databases like clinicaltrials.gov and ncbi.nlm.nih.gov/pubmed to double-check Watson’s findings.
“He did great,” Kolhe says, 100 percent on the work they double-checked. “One of the limitations with any computer or machine is it’s only as good as what you feed it. We feed him good information and let him go on that.”
Kolhe and Watson started testing on current patients in September and the 170-gene panel is now available to other pathologists. Kolhe and Illumina also have developed a more extensive 500-gene panel that Watson is currently digesting.
The pathologist thinks and hopes at some point Watson and the gene panels will become not only a common diagnostic tool for patients and their physicians, but a way to gauge treatment response and detect any additional mutations/variations the cancer is making in its parallel effort to survive.
“We are looking at that person and just his cancer,” Kolhe says.
“While this does not yet take us to the direct cause, because something caused the mutations, it takes us to the source – and hopefully to the best treatment – of his cancer.”
Editor’s note: Neither Kolhe nor the university will be compensated for their collaborations with IBM or Illumina unless patentable discoveries, like new cancer-causing genes, are made.
IBM Watson’s first commercial health care application came in 2013 to help with lung-cancer treatment decisions at Memorial Sloan Kettering Cancer Center in New York. He’s also helped forecast the weather and prepare tax returns.